Model Configuration and Assumptions
Starting from the operational equation of the blood-based Logan plot, Ichise et al. derived three multi-linear reference tissue model variants MRTM0, MRTM and MRTM2 [1]. They all assume an initial equilibration time t* from which on the derived multi-linear relation holds. However, if kinetics in the target tissue can be described by a 1-tissue compartment model (an assumption required for the SRTM), all data can be used for the fitting (t*=0). Otherwise an adequate t* value has to be determined.
Assuming the presence of receptor-devoid reference region TAC CT'(t), the target tissue TAC CT(t) is plotted as a function of the transformed tissue TACs as illustrated below. For the calculation of BPND it is assumed that the non-displaceable distribution volumes in the tissue and reference regions are identical.
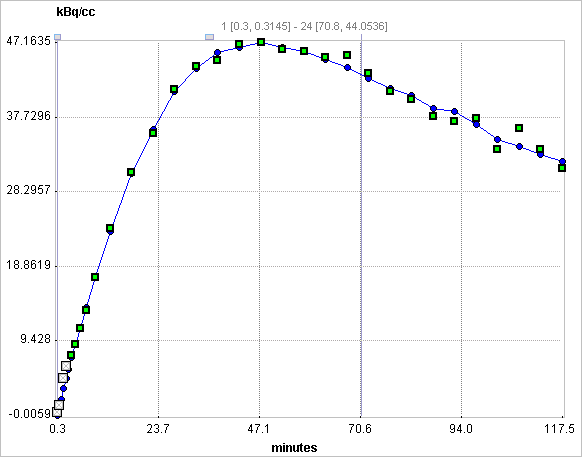
Operational Model Curve
To reduce noise-related bias effects arising in the MRTM0 method Ichise et al. applied a strategy known to be effective in reducing the noise-induced bias for the models requiring blood data. To this end the equation of the MRTM0 method was rearranged to remove the noisy tissue radioactivity term CT(t) from the independent variables. This approach resulted in a new method called MRTM with following operational equation for CT(t):
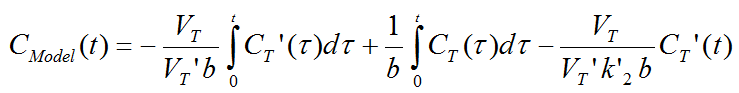
The multi-linear relationship above can be fitted using multi-linear regression, yielding three regression coefficients. The binding potential can then be calculated by dividing the first two regression coefficients
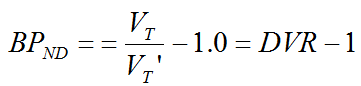
Furthermore, division of the first by the third regression coefficient yields an estimate of k2' .
Parameter Fitting
After switching to the Ichise NonInvasive MRTM model a suitable reference region must be selected. It allows fitting the multi-linear operational equation to the tissue TAC using the data segment starting from t*. The results are the three regression coefficients, and the binding potential BPnd and k2'.
A reasonable value of t* can be estimated by the based on the error criterion Max Err. For instance, if Max Err. is set to 10% and the fit box of t* is checked, the model searches the earliest sample such that the deviation between the regression and all measurements is less than 10%. Samples earlier than the t* time are disregarded for regression and thus painted in gray. In order to apply the analysis to the same data segment in all regions, please switch off the fit box of t*, propagate the model with the Copy to all Regions button, and then activate Fit all regions.
For receptor ligands with 1-tissue kinetics such as [11C]DASB the multi-linear equation is correct from t*=0, and the clearance rate constant from the tissue to plasma k2 is equal to the negative value of the second regression coefficient, -(1/b). Furthermore, R1 = K1/K'1, the relative radioligand delivery, equals the third regression coefficient.
Note: The reference methods MRTM2 and SRTM2 require k'2 as an input parameter. The k2' resulting from the MRTM method above might be a suitable estimate. Therefore, when switching in PKIN from the MRTM model to MRTM2 or SRTM2, k2' is automatically copied from MRTM, if the Model conversion option in the Extras panel is enabled.
Reference
1.Ichise M, Liow JS, Lu JQ, Takano A, Model K, Toyama H, Suhara T, Suzuki K, Innis RB, Carson RE: Linearized reference tissue parametric imaging methods: application to [11C]DASB positron emission tomography studies of the serotonin transporter in human brain. J Cereb Blood Flow Metab 2003, 23(9):1096-1112. DOI