The Batch Mode serves for the sequential processing of multiple data sets. The processing, which can be applied to the data sets, depends on their format:
▪KM file: A .km file saved from PKIN includes a complete definition of the blood interpolation model, the tissue model, weighting etc. for each region. Therefore, multiple processing types are supported including coupled fitting and Monte Carlo simulations. Additionally, during the fitting, the options on the Extras pane such as parameter initialization and randomized fitting will be effective. Note that the input file will be replaced by a new file with the results available in the fitting history.
▪Composite data file: A .kmData file contains only the data definition, except for the global specification of a model. Therefore, only the individual tissue model fitting is supported. The result will be saved as a new .km file with the same name as the input file, and with the results in the history.
Batch Facilities for .km Files
The Batch Mode entry from the Kinetic menu opens a dialog window for setting up the batch queue.
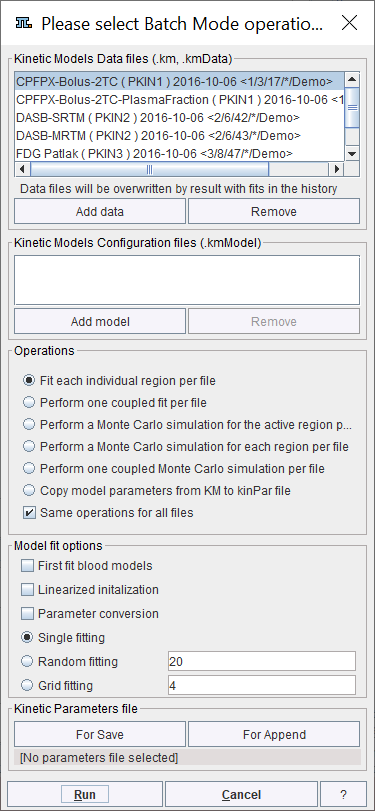
Use the Add data button for opening a data selection window and select the data sets to be processed from a database or the file system. Further data can incrementally be added, and selected list entries removed.
If only .km files have been selected it is assumed that appropriate configurations are included for the user's processing choice which is set by the radio button in the Operations section:
Fit each individual region |
All regional TACs are fitted with the respective models as defined in the .km file. |
Perform one coupled fit |
Performs coupled fitting with the model and coupling as defined in the .km file. |
Perform a Monte Carlo simulation for the active region |
Using the noise and model definitions in the file a Monte Carlo simulation of the current TAC is performed. |
Perform a Monte Carlo simulation for each region |
As above, but applied to all regional TACs. Can be used to evaluate the sensitivity at different parameter combinations. |
Perform one coupled Monte Carlo simulation |
Monte Carlo simulation using coupled fitting as defined in the file. |
Copy model parameters from KM to kinPar file |
Just a convenience to summarize the parameters of a series of files into one tabular text file. Note that a result file must first be defined with the For Save button for the option to become available. |
Fitting Options
The Model fit options give access to the mechanisms for improving objectivity and reliability of the results discussed in Fitting Options on Extras Panel with one exception. The First fit blood models option is intended for cases, where interpolation functions are defined for the different blood components but not yet fitted. If
Batch with Model Configuration Files
The batch mode also allows fitting multiple models to each data file. To do so, corresponding model configuration files can be selected with the Add model button in the Kinetic .
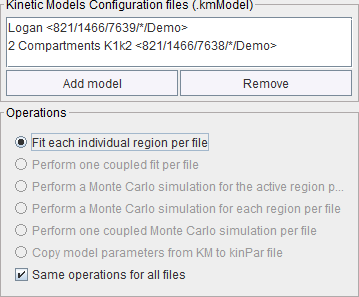
In this situation, only the Fit each individual region option is available. The results of the different model fits are saved in the regional model histories.
Batch Results
The For Save and the For Append buttons can be used to specify a file as a container for saving the result parameters of the batch run. These results are in the .kinPar and can be transferred to the R console for performing statistics.
For allowing visual inspection of the results each original data file is replaced by new file. It contains in the history all model fits from the batch, so that a user can simply load the file and step through the fits using the history buttons.
Starting the Batch
The Run button starts processing, and PMOD will be blocked until processing completes.